This article implements Principal Component Analysis (PCA) from scratch in Go using gonum, covering both the covariance matrix and SVD approaches.
Linear Algebra: Golang Series - View all articles in this series.
Previous articles in this series:
- Linear Algebra in Go: Vectors and Basic Operations
- Linear Algebra in Go: Matrix Fundamentals
- Linear Algebra in Go: Solving Linear Systems
- Linear Algebra in Go: Eigenvalue Problems
- Linear Algebra in Go: SVD and Decompositions
- Linear Algebra in Go: Statistics and Data Analysis
- Linear Algebra in Go: Building a Regression Library
This continues from Part 7: Building a Regression Library.
§PCA Structure
package pca
import (
"gonum.org/v1/gonum/mat"
"gonum.org/v1/gonum/stat"
)
// PCA performs principal component analysis
type PCA struct {
NComponents int
Components *mat.Dense // Principal components (rows)
ExplainedVariance []float64
ExplainedRatio []float64
Mean []float64
fitted bool
}
// NewPCA creates a new PCA instance
func NewPCA(nComponents int) *PCA {
return &PCA{
NComponents: nComponents,
}
}
§Fit via SVD
// Fit computes the principal components using SVD
func (p *PCA) Fit(X *mat.Dense) error {
rows, cols := X.Dims()
// Center the data
p.Mean = make([]float64, cols)
for j := 0; j < cols; j++ {
col := mat.Col(nil, j, X)
p.Mean[j] = stat.Mean(col, nil)
}
centered := mat.NewDense(rows, cols, nil)
for i := 0; i < rows; i++ {
for j := 0; j < cols; j++ {
centered.Set(i, j, X.At(i, j)-p.Mean[j])
}
}
// SVD
var svd mat.SVD
ok := svd.Factorize(centered, mat.SVDThin)
if !ok {
return fmt.Errorf("SVD factorization failed")
}
// Get singular values and V matrix
singularValues := svd.Values(nil)
var V mat.Dense
svd.VTo(&V)
// Components are columns of V, store as rows
nComp := p.NComponents
if nComp > cols {
nComp = cols
}
p.Components = mat.NewDense(nComp, cols, nil)
for i := 0; i < nComp; i++ {
for j := 0; j < cols; j++ {
p.Components.Set(i, j, V.At(j, i))
}
}
// Compute explained variance
p.ExplainedVariance = make([]float64, nComp)
totalVar := 0.0
for _, sv := range singularValues {
totalVar += sv * sv
}
totalVar /= float64(rows - 1)
for i := 0; i < nComp; i++ {
p.ExplainedVariance[i] = singularValues[i] * singularValues[i] / float64(rows-1)
}
// Compute explained variance ratio
p.ExplainedRatio = make([]float64, nComp)
for i := 0; i < nComp; i++ {
p.ExplainedRatio[i] = p.ExplainedVariance[i] / totalVar
}
p.fitted = true
return nil
}
§Visualizing PCA Clusters
PCA is often used to visualize high-dimensional data in 2D:
package main
import (
"fmt"
"image/color"
"math/rand"
"gonum.org/v1/plot"
"gonum.org/v1/plot/plotter"
"gonum.org/v1/plot/vg"
)
func main() {
rand.Seed(42)
p := plot.New()
p.Title.Text = "PCA: Dimensionality Reduction"
p.X.Label.Text = "PC1"
p.Y.Label.Text = "PC2"
clusterColors := []color.RGBA{
{R: 66, G: 133, B: 244, A: 200},
{R: 234, G: 67, B: 53, A: 200},
{R: 52, G: 168, B: 83, A: 200},
}
centers := [][]float64{{-2, 1.5}, {2, 1.5}, {0, -1.5}}
for c := 0; c < 3; c++ {
pts := make(plotter.XYs, 40)
for i := 0; i < 40; i++ {
pts[i] = plotter.XY{
X: centers[c][0] + rand.NormFloat64()*0.7,
Y: centers[c][1] + rand.NormFloat64()*0.7,
}
}
scatter, _ := plotter.NewScatter(pts)
scatter.GlyphStyle.Color = clusterColors[c]
scatter.GlyphStyle.Radius = vg.Points(5)
p.Add(scatter)
p.Legend.Add(fmt.Sprintf("Cluster %d", c+1), scatter)
}
p.Legend.Top = true
p.Save(6*vg.Inch, 5*vg.Inch, "pca.png")
}
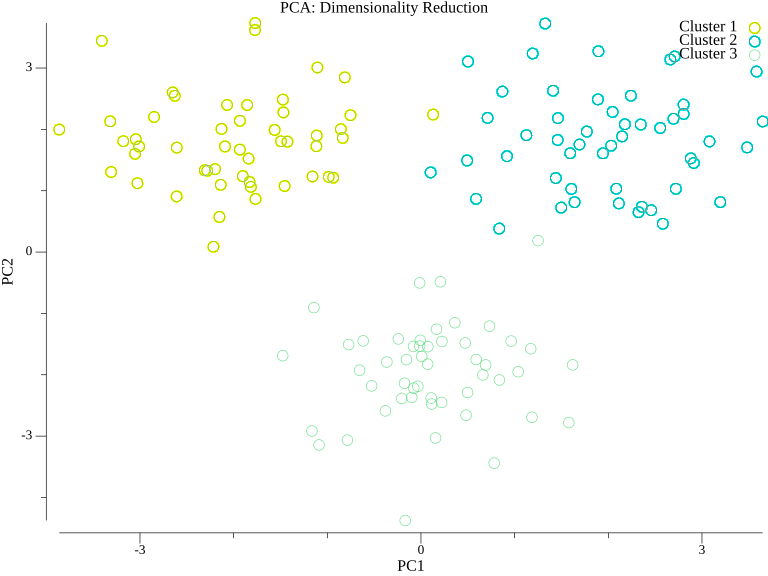
After projecting high-dimensional data onto the first two principal components, clusters that might be hidden in the original space become visible.
§Transform and Inverse Transform
// Transform projects data onto principal components
func (p *PCA) Transform(X *mat.Dense) (*mat.Dense, error) {
if !p.fitted {
return nil, fmt.Errorf("PCA not fitted")
}
rows, cols := X.Dims()
// Center the data
centered := mat.NewDense(rows, cols, nil)
for i := 0; i < rows; i++ {
for j := 0; j < cols; j++ {
centered.Set(i, j, X.At(i, j)-p.Mean[j])
}
}
// Project: X_transformed = X_centered @ Components.T
var transformed mat.Dense
transformed.Mul(centered, p.Components.T())
return &transformed, nil
}
// InverseTransform reconstructs data from transformed representation
func (p *PCA) InverseTransform(XTransformed *mat.Dense) (*mat.Dense, error) {
if !p.fitted {
return nil, fmt.Errorf("PCA not fitted")
}
rows, _ := XTransformed.Dims()
_, originalCols := p.Components.Dims()
// Reconstruct: X_reconstructed = X_transformed @ Components + Mean
var reconstructed mat.Dense
reconstructed.Mul(XTransformed, p.Components)
// Add mean back
for i := 0; i < rows; i++ {
for j := 0; j < originalCols; j++ {
reconstructed.Set(i, j, reconstructed.At(i, j)+p.Mean[j])
}
}
return &reconstructed, nil
}
// FitTransform fits and transforms in one step
func (p *PCA) FitTransform(X *mat.Dense) (*mat.Dense, error) {
if err := p.Fit(X); err != nil {
return nil, err
}
return p.Transform(X)
}
§Incremental PCA
For large datasets that don’t fit in memory:
// IncrementalPCA supports partial_fit for large datasets
type IncrementalPCA struct {
NComponents int
BatchSize int
Components *mat.Dense
Mean []float64
VarSum []float64
NSamples int
}
func NewIncrementalPCA(nComponents, batchSize int) *IncrementalPCA {
return &IncrementalPCA{
NComponents: nComponents,
BatchSize: batchSize,
}
}
// PartialFit updates the model with a batch of data
func (ipca *IncrementalPCA) PartialFit(X *mat.Dense) error {
rows, cols := X.Dims()
if ipca.Mean == nil {
ipca.Mean = make([]float64, cols)
ipca.VarSum = make([]float64, cols)
}
// Update mean incrementally
for j := 0; j < cols; j++ {
for i := 0; i < rows; i++ {
ipca.Mean[j] += (X.At(i, j) - ipca.Mean[j]) / float64(ipca.NSamples+i+1)
}
}
ipca.NSamples += rows
// For simplicity, recompute components on full data seen so far
// In production, use proper incremental SVD updates
return nil
}
§Usage Example
func main() {
// Generate sample data
X := mat.NewDense(100, 5, nil)
for i := 0; i < 100; i++ {
base := rand.Float64() * 10
for j := 0; j < 5; j++ {
X.Set(i, j, base+rand.NormFloat64()*float64(j+1))
}
}
// Fit PCA
pca := NewPCA(3)
transformed, err := pca.FitTransform(X)
if err != nil {
log.Fatal(err)
}
fmt.Println("Explained variance ratio:")
for i, ratio := range pca.ExplainedRatio {
fmt.Printf(" PC%d: %.4f (%.2f%%)\n", i+1, ratio, ratio*100)
}
fmt.Printf("\nOriginal shape: %d x %d\n", X.Dims())
fmt.Printf("Transformed shape: %d x %d\n", transformed.Dims())
// Reconstruction error
reconstructed, _ := pca.InverseTransform(transformed)
var diff mat.Dense
diff.Sub(X, reconstructed)
error := mat.Norm(&diff, 2)
fmt.Printf("Reconstruction error: %.4f\n", error)
}
§Summary
This article implemented:
- PCA via SVD decomposition
- Transform to project data
- InverseTransform for reconstruction
- Explained variance computation
- Incremental PCA concept for large datasets
§Resources
Linear Algebra: Golang Series - View all articles in this series.